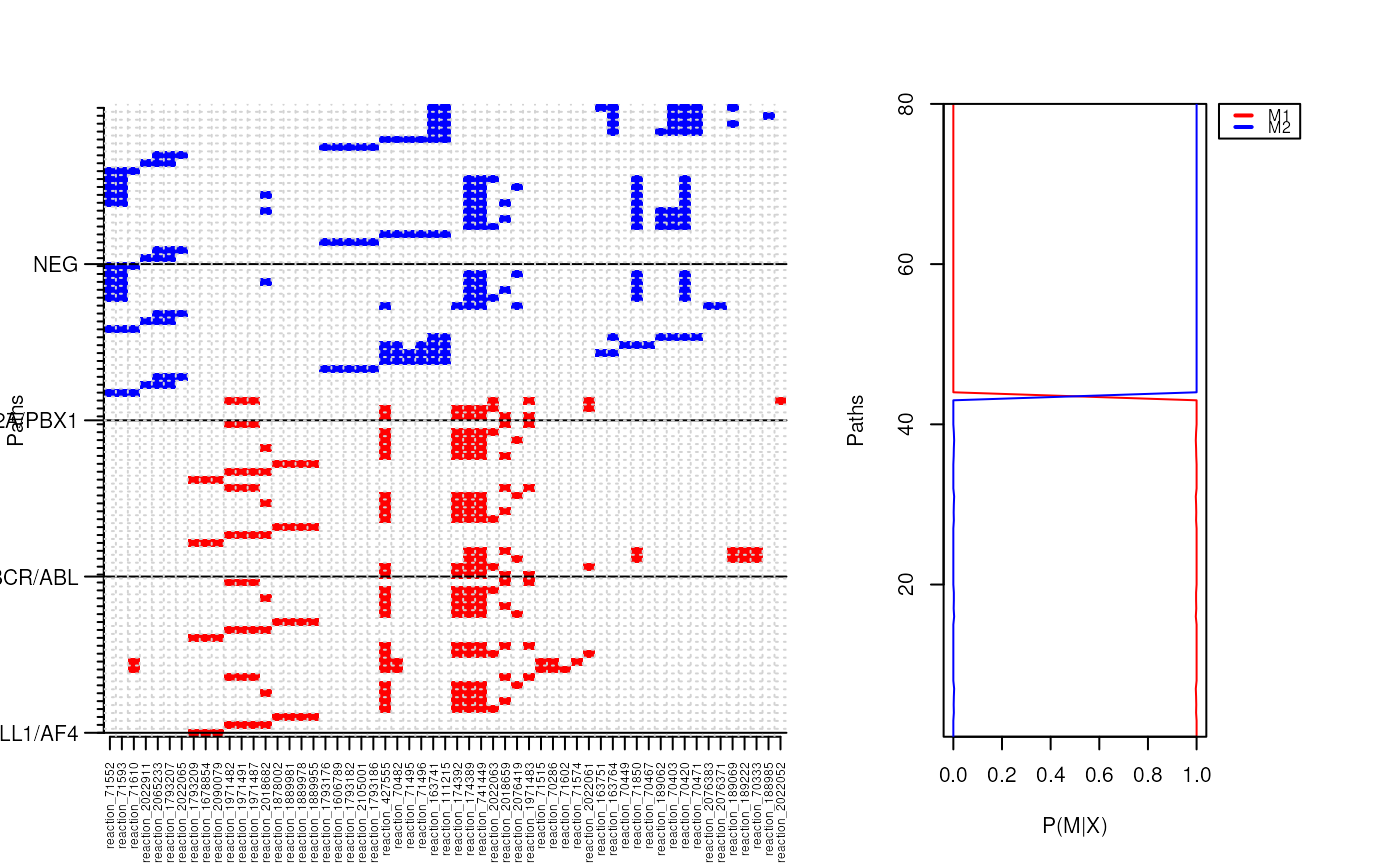
Plots the structure of all path clusters
Arguments
- ybinpaths
The training paths computed by
pathsToBinary.- clusters
The pathway cluster model trained by
pathClusterorpathClassifier.- col
Colors for each path cluster.
- grid
A logical, whether to add a
gridto the plot- ...
Extra paramaters passed to
plotClusterMatrix
Value
plotClusterMatrix plots an image of all paths the training dataset. Rows are the paths and columns
are the genes (features) included within each path. Paths are colored according to cluster membership.
plotClusterProbs The training set posterior probabilities for each path belonging to a 3M component.
plotClusters: combines the two plots produced by plotClusterProbs and plotClusterMatrix.
See also
Other Path clustering & classification methods:
pathClassifier(),
pathCluster(),
pathsToBinary(),
plotClassifierROC(),
plotPathClassifier(),
plotPathCluster(),
predictPathClassifier(),
predictPathCluster()
Other Plotting methods:
colorVertexByAttr(),
layoutVertexByAttr(),
plotAllNetworks(),
plotClassifierROC(),
plotCytoscapeGML(),
plotNetwork(),
plotPathClassifier(),
plotPaths()
Examples
## Prepare a weighted reaction network.
## Conver a metabolic network to a reaction network.
data(ex_sbml) # bipartite metabolic network of Carbohydrate metabolism.
rgraph <- makeReactionNetwork(ex_sbml, simplify=TRUE)
#> This graph was created by an old(er) igraph version.
#> ℹ Call `igraph::upgrade_graph()` on it to use with the current igraph version.
#> For now we convert it on the fly...
## Assign edge weights based on Affymetrix attributes and microarray dataset.
# Calculate Pearson's correlation.
data(ex_microarray) # Part of ALL dataset.
rgraph <- assignEdgeWeights(microarray = ex_microarray, graph = rgraph,
weight.method = "cor", use.attr="miriam.uniprot",
y=factor(colnames(ex_microarray)), bootstrap = FALSE)
#> 100 genes were present in the microarray, but not represented in the network.
#> 55 genes were couldn't be found in microarray.
#> Assigning edge weights for label ALL1/AF4
#> Assigning edge weights for label BCR/ABL
#> Assigning edge weights for label E2A/PBX1
#> Assigning edge weights for label NEG
## Get ranked paths using probabilistic shortest paths.
ranked.p <- pathRanker(rgraph, method="prob.shortest.path",
K=20, minPathSize=8)
#> Extracting the 20 most probable paths for ALL1/AF4
#> Extracting the 20 most probable paths for BCR/ABL
#> Extracting the 20 most probable paths for E2A/PBX1
#> Extracting the 20 most probable paths for NEG
## Convert paths to binary matrix.
ybinpaths <- pathsToBinary(ranked.p)
p.cluster <- pathCluster(ybinpaths, M=2)
plotClusters(ybinpaths, p.cluster, col=c("red", "blue") )