Converts the result from pathRanker into something suitable for pathClassifier or pathCluster.
Source:R/pathClassifier.R
pathsToBinary.RdConverts the result from pathRanker into something suitable for pathClassifier or pathCluster.
pathsToBinary(ypaths)Arguments
- ypaths
The result of
pathRanker.
Value
A list with the following elements.
- paths
All paths within ypaths converted to a binary string and concatenated into the one matrix.
- y
The response variable.
- pidx
An matrix where each row specifies the location of that path within the
ypathsobject.
Details
Converts a set of pathways from pathRanker
into a list of binary pathway matrices. If the pathways are grouped by a response label then the
pathsToBinary returns a list labeled by response class where each element is the binary
pathway matrix for each class. If the pathways are from pathRanker then a list wiht
a single element containing the binary pathway matrix is returned. To look up the structure of a
specific binary path in the corresponding ypaths object simply use matrix index by calling
ypaths[[ybinpaths\$pidx[i,]]], where i is the row in the binary paths object you
wish to reference.
See also
Other Path clustering & classification methods:
pathClassifier(),
pathCluster(),
plotClassifierROC(),
plotClusterMatrix(),
plotPathClassifier(),
plotPathCluster(),
predictPathClassifier(),
predictPathCluster()
Examples
## Prepare a weighted reaction network.
## Conver a metabolic network to a reaction network.
data(ex_sbml) # bipartite metabolic network of Carbohydrate metabolism.
rgraph <- makeReactionNetwork(ex_sbml, simplify=TRUE)
#> This graph was created by an old(er) igraph version.
#> ℹ Call `igraph::upgrade_graph()` on it to use with the current igraph version.
#> For now we convert it on the fly...
## Assign edge weights based on Affymetrix attributes and microarray dataset.
# Calculate Pearson's correlation.
data(ex_microarray) # Part of ALL dataset.
rgraph <- assignEdgeWeights(microarray = ex_microarray, graph = rgraph,
weight.method = "cor", use.attr="miriam.uniprot",
y=factor(colnames(ex_microarray)), bootstrap = FALSE)
#> 100 genes were present in the microarray, but not represented in the network.
#> 55 genes were couldn't be found in microarray.
#> Assigning edge weights for label ALL1/AF4
#> Assigning edge weights for label BCR/ABL
#> Assigning edge weights for label E2A/PBX1
#> Assigning edge weights for label NEG
## Get ranked paths using probabilistic shortest paths.
ranked.p <- pathRanker(rgraph, method="prob.shortest.path",
K=20, minPathSize=6)
#> Extracting the 20 most probable paths for ALL1/AF4
#> Extracting the 20 most probable paths for BCR/ABL
#> Extracting the 20 most probable paths for E2A/PBX1
#> Extracting the 20 most probable paths for NEG
## Convert paths to binary matrix.
ybinpaths <- pathsToBinary(ranked.p)
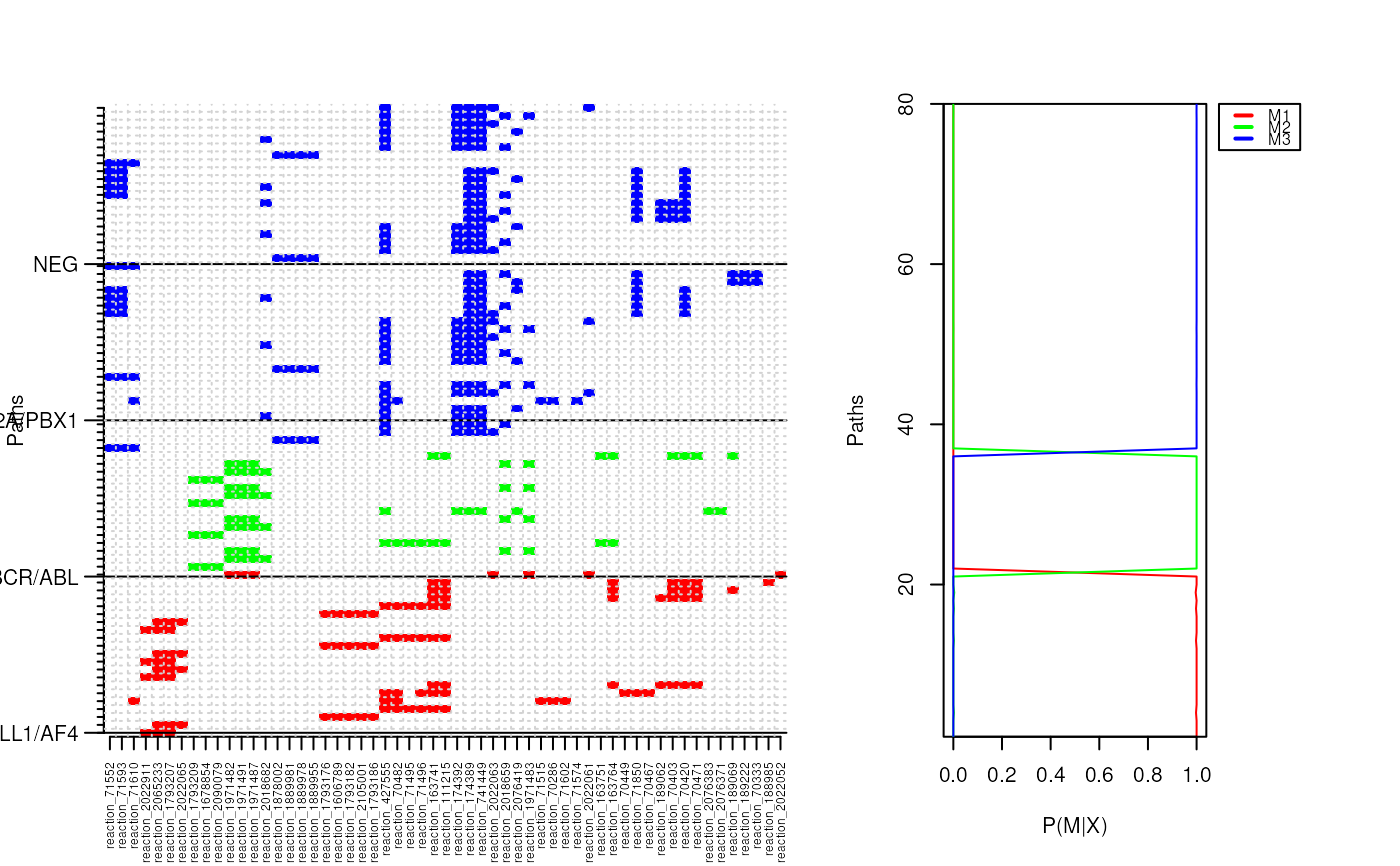
p.cluster <- pathCluster(ybinpaths, M=3)
plotClusters(ybinpaths, p.cluster, col=c("red", "green", "blue") )